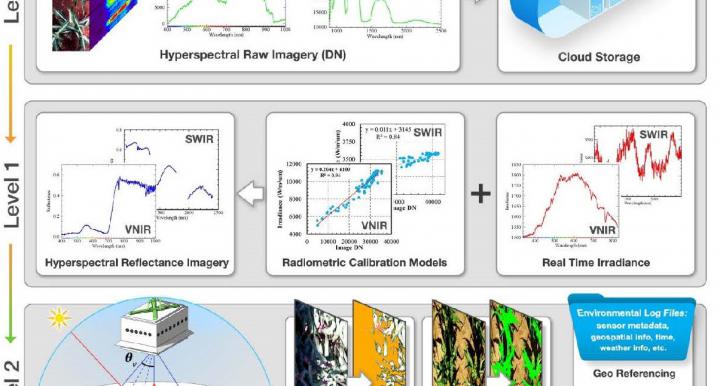
TERRA REF Crop Sensing Components
Breeding is currently limited by the speed at which phenotypes can be measured, and the information that can be extracted from these measurements. TERRA REF is generating reference data sets from a variety of sensors and sensing platforms.
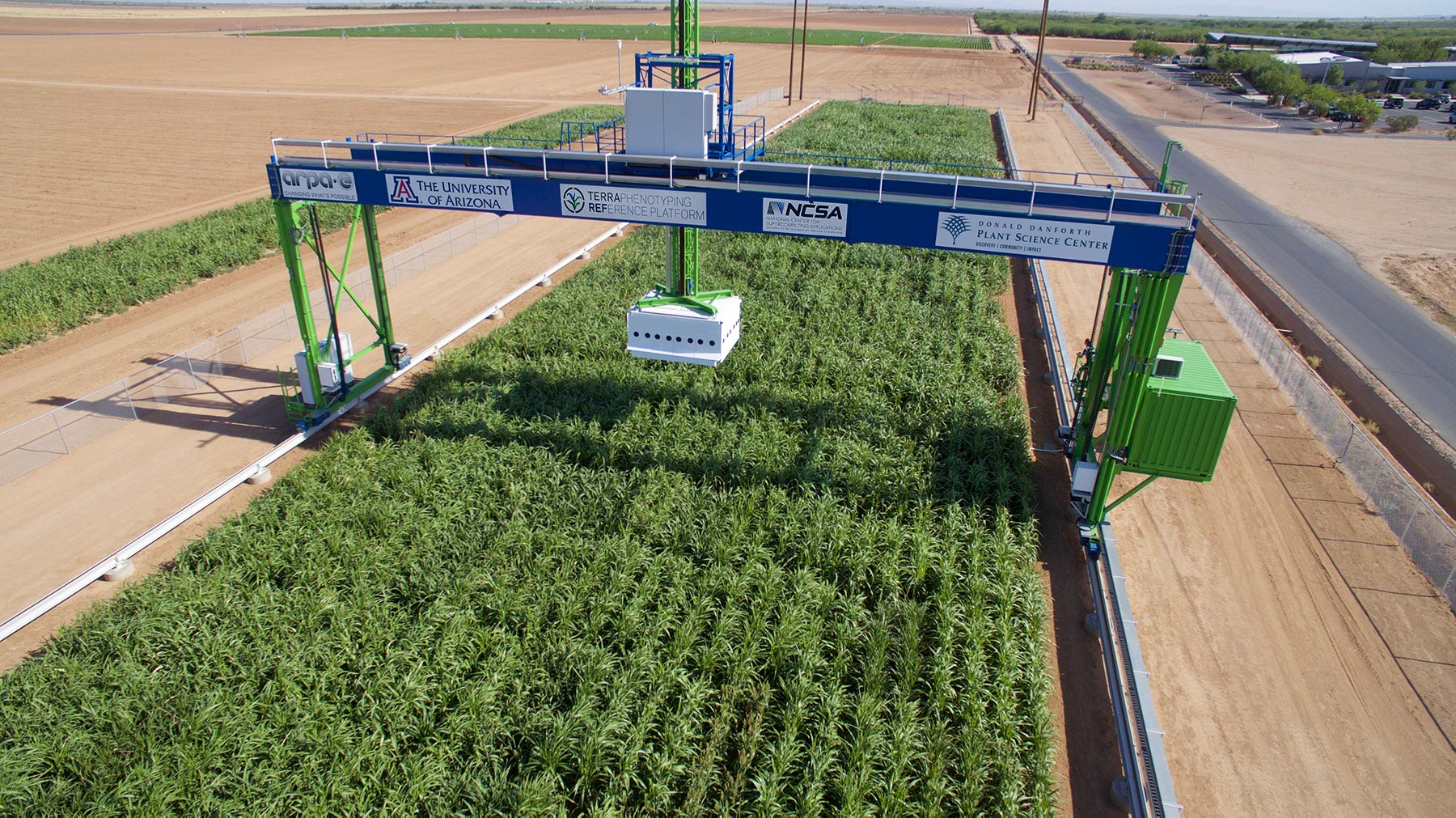
Lemnatec Field Scanner
The World's largest agricultural robot collects data at unprecedented spatial, temporal, and spectral resolution.
Controlled Environment Phenotyping
Indoor system measures over a thousand plants each day.
Unmanned Aerial Vehicles
Concurrent field scanning with diverse sensors deployed on fixed wing and rotary platforms
Ground Based Vehicles
Proximate sensing using tractors and push-carts.
Genomic and genetic data
Whole genome resequencing and genotyping-by sequencing data from 100s of Sorghum and Wheat accessions.
Open data, software, and computing
Fully open phenomics pipeline and data.